Generative models for networks with higher–order motifs
We base our approach on generative models where vertices are connected not only by single edges but also copies of higher order subgraphs, that we call “atoms”. Assuming such a generative model leads us to consider latent spaces that correspond the atomic subgraphs added to the graph during the generation process. We call these latent states subgraph configurations. Subgraph configurations are decompositions of a network into smaller subgraphs and can be thought of as generalized hypergraphs where hyper edges have the form of any connected motif. More formally a subgraph configuration C on a set of vertices V is a set of subgraphs of the complete graph KV on V. Given such a C we define the atoms of C, M(C), as the set of motifs occurring in C. For a graphical illustration of a subgraph configuration see Fig. 1. Note that any subgraph configuration C can be transformed into a graph G by simply taking the union of the edges contained in the subgraphs in C i.e. E(G) = ⋃S∈CE(S). We says a subgraph configuration C covers G if E(G) = ⋃S∈CE(S).
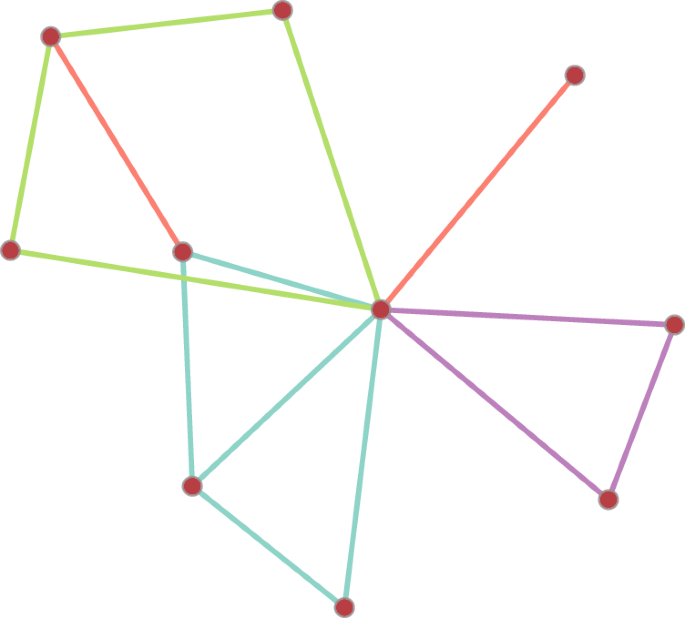
A subgraph configuration consisting two single edges, a triangle, a 4-cycle and a diamond. Edges are coloured according to the type of higher order subgraphs.
In the formulation of our method we use maximum entropy ensembles of subgraph configurations that results from imposing hard constraints on the counts and distributions of atomic subgraphs used to construct the network5. In these models, which are also known as microcanonical ensembles, all subgraph configurations that satisfy the given constraints are equi–probable. Consequently, the likelihood of configurations can be computed by simply counting the number of configurations that are compatible with the given constraints. Another advantage of using microcanonical models is that every configuration is compatible with a single set of parameters which allows marginal probabilities to be computed in closed form without computing costly integrals.
Motifs, automorphisms and orbits
First, we review some basic concepts that are essential in formulating the generative model starting with graph isomorphism. Two graphs G and H, are said to be isomorphic if there exists a bijection ϕ: V(G) → V(H) such that \((v,v\prime )\in E(G)\ \iff \ (\phi (v),\phi (v\prime ))\in E(H)\). Being isomorphic is an equivalence relation of which the equivalence classes correspond to unlabelled graphs also known as motifs. We denote motifs using lower-case letters and write G ≃ g. Automorphisms are special types of isomorphisms which map a graph to itself. Automorphisms are essentially vertex permutations that leave the structure of the graph unchanged. The set of all automorphisms of G form a group which we denote as Aut(G). Finally, the orbits of a graph are subsets of vertices that can be mapped onto each other by Aut(G) and correspond to classes of structurally identical vertices of G (see Fig. 2). The ith orbit of G is denoted as OG,i.
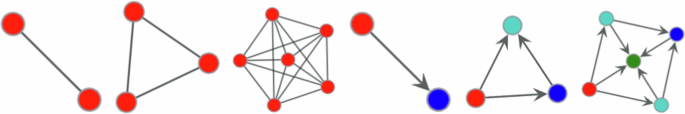
Some undirected and directed motifs with vertex colours indicating different orbits.
In the remainder of the text we shall not explicitly distinguish between undirected and directed graphs because in the context of subgraph configurations this distinction can be reduced to the set of atomic subgraphs and the character of their symmetries. The only modification required for the directed case is in the definition of graph isomorphisms which for directed graphs are required to also conserve edge directions.
Microcanonical models with fixed subgraph counts
The simplest type of subgraph configuration model (SGCM) on N vertices with atoms M = m is the ensemble of all subgraph configurations that contain exactly nm subgraphs of type m ∈ M. In the microcanonical ensemble all such configurations have equal probability which can be computed by simply counting the total number of such configurations:
$$P(C| \bfn_m,M)= \\ n_m\endarray\right)\right)^-1,$$
(1)
where \({{\mathcalH}}_N,m\) is set of all m-subgraphs of the complete graph KN. It follows from the definition of the automorphism group that:
$$| {{{{{\mathcalH}}}}}_N,m| =\fracN! Aut(m).$$
(2)
In such models atomic subgraphs are distributed uniformly over the vertices and hence we shall refer to these types of models as homogeneous SGCMs. When M contains only the single edge motif, homogeneous models reduce to the classical Erdös Rényi random graph GN,e where all graphs with N vertices and e edges are equi–probable. Throughout this article we assume that networks are sparse i.e. nm = O(N) for which \(\log (P)\) in Eq. (1) can be approximated using Stirling’s formula24.
Subgraph configuration models and random graphs
Distributions over subgraph configurations such as the one introduced above can be turned into distributions over graphs by projecting subgraph configurations onto graphs. In general we shall assume that the graphs under consideration are simple i.e. that they do not contain parallel edges or self loops. In this case subgraph configurations can be projected onto graphs by simply taking the union of the edges of the subgraphs in a given configuration C. Using this projection any distribution over subgraph configurations C, denoted P(C), can be mapped onto a distribution over graphs where the probability of graph G is given by the sum of the probabilities of the subgraph configurations of which the projection is G, i.e.:
$$P(G)=\sum\limits_ \bigcup _s\in CE(s)=E(G)P(C).$$
(3)
Note that subgraph configurations differ from other types of latent states, such as community assignments in the SBM, in that each subgraph configuration maps to a unique graph. Therefore, subgraph configurations can be seen as a general class of exact/lossless graph representations that includes many known graph representations including edge lists, adjacency list and bipartite representations.
Degree–corrected models
In homogeneous SGCMs atomic subgraphs are distributed uniformly over the vertices which in turn results in graphs with Poisson type degree distributions3. Therefore homogeneous models are in general not a good fit for empirical networks that have highly heterogeneous or power-law type degree distributions. Thus we consider degree–corrected subgraph configuration models where one not only constrains the counts of atomic subgraphs but also the number of atomic subgraphs attached to each vertex. For this we consider orbit degree sequences of configurations where dm,i(C)(v) specifies the number m-subgraphs in C that are attached to v at orbit i. For instance the central vertex of the configuration in Fig. 1 has edge degree 1, triangle degree 1, 4-cycle degree 1. Whereas these motifs all have a single orbit the diamond has two orbits i.e. one corresponding to the vertices with degree 2 and one corresponding to the vertices with degree 3 for which the central vertex of the configuration in Fig. 1 has degrees 0 and 1, respectively. Note that the orbit degree sequence is defined with respect to a subgraph configuration C and should not be confused with the number of subgraphs of type m containing a certain vertex v in the graph. As in the case of fixed subgraph counts the microcanonical ensemble of subgraph configurations having orbit degree sequence dm,i(v) is defined as the uniform ensemble over all configurations having orbit degree sequence dm,i(C)(v). As a result the probability of configurations can be calculated by counting the number of configurations with orbit degree sequence dm,i(C)(v)5. A brief derivation of the likelihood can be found in the Methods section. The degree–corrected subgraph configuration model is closely related to the model by Karrer and Newman introduced in4 which is defined in terms of a stub-matching process similar to the (edge) configuration model.
Note that the degree of a vertex v (counting parallel edges) is fully determined by the orbit degree:
$$d(v)=\sum\limits_m,id_m,i(v)d(O_m,i),$$
(4)
where d(Om,i) is the degree of vertices in orbit Om,i and hence the degree corrected model can model networks with heterogeneous degree distributions. Although we discard parallel-edges in our formulation the expected number of such multi-edges is O(1) for sparse models.
Coarse grained degree corrected models
Constraining the atomic degrees at the level of orbits in certain cases significantly increases the model complexity of DC-SGCMs due to the fact that the model requires a degree sequence of length ∣V∣ to be specified for every orbit of each atom in M. Although the Bayesian approach safeguards against over–fitting by balancing goodness of fit and parametric complexity, using models with high parametric complexity can inhibit the detection of regularities, i.e. lead to under-fitting, due to the high complexity cost associated to including such regularities in the model. Thus, in the context of statistical inference degree–corrected models that have lower model complexity for the same set of atoms need to be considered. It should however be iterated that the final choice should be made on the basis of how well each model fits the data, for instance as we shall do in this paper, via the posterior odds ratio.
The parametric complexity of DC-SGCMs can be reduced by aggregating components of the orbit degree sequence at different levels of granularity. One such option is to constrain atomic degrees at the level of motifs i.e. by the fixing the sum dm(v) = ∑idm,i(v) instead of each dm,i(v) separately which results in a model that requires ∣M∣ degree sequences. The parametric complexity of the model can be even further reduced by only constraining the total number of atomic subgraphs attached to each vertex i.e. dt(v) = ∑m∑idm,i(v) which results in a model that only requires a single degree sequence. In the case of directed graphs we also consider a model where orbits are grouped according to the their in– and out–degrees because grouping orbits according to atoms will in general lead to a mixing of in– and out–degrees which in turn results in models where edge directions are random. For this we group orbits in to three groups corresponding to orbits that only have incoming edges (\(d_in(v)=\sum _m,id_m,i\)), only out going edges and have both in– and out–going edges, respectively. In this model the atomic degree sequence has 3 components and hence can be seen as a generalization of the configuration model where the in–, out– and mutual–edge degrees are conserved separately1. The likelihood of coarse grained ensembles can be obtained in closed form by the same procedure as the orbit degree model5.
Inference
Our goal is to infer a subgraph configuration C for given the graph G. Hence, we consider P(C∣G) which can be obtained via Bayes’ theorem:
$$P(C| G)=\fracP(GP(G),$$
(5)
where P(G∣C) = 1 if ⋃s∈CE(s) = E(G) and 0 otherwise, and P(C) is given by the prior over subgraph configurations. Note that subgraph configurations are latent structures that fully determine the graph and therefore differ from other latent spaces such as node partitions in the SBM that do not uniquely determine the graph.
We first consider the case of degree-corrected subgraph configuration models. For an atomic degree sequence dm,o having motif set M and count vectors nm we assume a nested prior with a conditional dependence structure of the following form:
$$P(C)=P(C| \bfd_\bfm,\bfi)P(\bfd_{\bfm,{{{\bfi}}}}| \bfn_\bfm)P(\bfn_\bfm| M,e(G))P(M),$$
(6)
where e(G) is the number of edges in G. The above form applies to all variants of the degree–corrected SGCM, the only difference being the number of components in the degree sequence. Similarly, in the case of the homogeneous SGCMs we consider a prior that has the following form:
$$P(C)=P(C| \bfn_\bfm)P(\bfn_{\bfm}| M,e(G))P(M).$$
We seek to identify a subgraph configuration with maximum posterior probability (MAP) and will call such a configuration a MAP-configuration. Note that this is equivalent to inferring the latent state of the model rather than the parameters of the model. However, the two problems are closely related through Eq. (3).
Model selection
The model that is most suitable for describing a given network can be selected via posterior odds ratio. For this we first find the MAP-configuration for each model variant of the subgraph configuration model including homogeneous models. In the degree corrected case we consider models that constrain the distributions of atomic subgraphs at the level of orbits, atoms and total number of subgraphs. For directed networks in addition to these we also consider the directed orbit degree model. We then select the model-configuration combination that has highest posterior probability, which in turn is equivalent to selecting the configuration with the shortest description length. A more detail explanation of the model selection procedure is given in the methods section.
Minimum description length
The Bayesian formulation outlined above is equivalent to finding a subgraph configuration that has Minimum Description Length (MDL)25. Although the equivalence of MDL and Bayesian approaches holds more broadly, in our case it is more directly evident due to the discrete nature of the parameters.
$$\Sigma (C)=S(C| \bfd,{{{{{\bfn}}}}}_{{{{{\bfm}}}}},M)+\epsilon (\bfd,{{{{{\bfn}}}}}_{{{{{\bfm}}}}},M),$$
(7)
where Σ(C) is the description length (DL), \(S(C)=-\log (P(C| {{{{\bfd}}}},{{{{{\bfn}}}}}_{{{{{\bfm}}}}},M))\) the entropy which corresponds to the information required to specify the positions of the subgraphs given the model parameters and \(\epsilon ({{{{\bfd}}}},{{{{{\bfn}}}}}_{{{{{\bfm}}}}},M)=-\log (P({{{{\bfd}}}},{{{{{\bfn}}}}}_{{{{{\bfm}}}}},M))\) is the information required to describe the parameters of the model i.e. the model complexity. Consequently, finding a MAP-configuration is equivalent to finding a configuration with minimum description length.
The atomic structure of real world networks
Due to the computational complexity of the problem we restrict the set of motifs included in the analysis to connected motifs up to size 5 for directed networks and to motifs up to size 8 for undirected networks. There are 9578 such motifs in the directed case and 12112 in the undirected case. Despite the large number of potential motifs included in the analysis our method is able to identify concise sets of atoms (see ∣Mopt∣ in Table 1) that cover a large proportion of the edges in each of these networks.
Model selection
In this section we compare different subgraph configuration models, and their fit to data. For this, we consider different models introduced in the text so far, namely the homogeneous micro-canonical ensemble, the orbit degree–corrected model, the atomic degree model and the total degree model. For directed networks we also consider directed degree model where orbits are aggregated based on edge directions. The MAP configurations corresponding to different types of degree–corrected models tend to be quite similar both in terms of the atoms and subgraphs they contain. In general though, the number of atoms in the MAP configuration tends to increase as the level of detail at which the atomic degree sequence is conserved decreases since this decreases the model complexity cost of incorporating new atoms into the configuration.
Once we obtain the MAP-configuration for each model we select the optimal configuration-model pair by comparing their posterior odds ratio/description lengths, see Table 1. We find that for undirected networks the total degree model results in the shortest description length, except for the Malaria Genes network. For the directed networks we find that in general the directed degree model or total degree model result in the shortest description lengths.
Our results show that the inclusion of higher order interactions in representations of real world networks leads to much more parsimonious representations when compared to their counterparts that only include dyadic interactions (i.e. single edges) with reductions in DL ranging from 297 to 13,007 nats. Note that this reduction in description length directly relates to the posterior odds ratio of the MAP configuration CM and the configuration consisting of all single edges i.e. E(G) :
$$\fracP(C_MP(E(G)=\exp (\Sigma (E(G))-\Sigma (C_M)),$$
where Σ(E(G)) is the DL of E(G) as given by the edge only configuration model. In other words we find that the networks in Table 1 are exponentially more likely to have been generated by the higher order models corresponding to their MAP configurations than by the edge only configuration model. Similarly, for real world networks we analysed, we find that degree corrected models in general result in significantly shorter description lengths than non–degree corrected models indicating that degree corrected models are an overall better fit to the data.
In the following section we provide a more detailed discussion of the MAP configurations for some of the networks in Table 1. The results for the remaining networks are discussed in Supplementary Note 5.
Co-authorship network of network scientists
We first consider a widely studied co-authorship network between network scientists26. We find that the MAP configuration (see Figure 3) of this network contains 15 nontrivial atoms other than the single edge that cover over 85% of the edges (see Table 2). As expected in a collaboration network the MAP configuration contains many cliques corresponding to publications with more than two authors. In addition to cliques we also find higher order interactions where two high degree coauthors collaborate with the same group of researchers (Table 2 Atoms 3 and 12) as well as patterns where two high degree authors collaborate with the same group of lower degree nodes but not with each other (Table 2 Atoms 5,7,8,13 and 14).
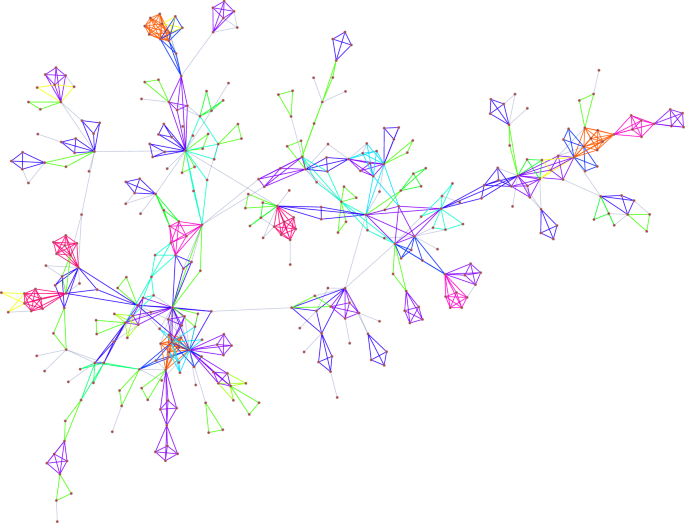
Visualization of the largest connected component of the collaboration network between network scientists. Colours represent different atomic subgraph types in the MAP-configuration. A high resolution plot of the full network can be found in Supplementary Fig. 1.
Human connectome
Next we consider a directed connectome of the human brain from ref. 27. The network has 1015 vertices and 4,008 edges. We find that the MAP configuration contains 20 non-trivial patterns which cover ~75% of the edges. The MAP configuration contains a large number of feed-forward loops (FFL) (Atom 1 in Table 3) and bi–fan motifs (Atom 5 in Table 3) usually associated with neuronal networks. We also find a large number of acyclic square shaped motifs (Atoms 2 and 3 in Table 3) that can be interpreted as 4-node generalizations of the FFL, where an input node is connected to an output node via both direct connections and indirect connections in the form of longer directed paths. In addition to these basic types we also recover more complex atoms that can be interpreted as various combinations of these lower order atoms. Such larger arrangements of smaller motifs have recently been studied in ref. 28. However, our result indicate that in the connectome lower order motifs combine in denser and more complex patterns than the pairwise combinations of lower order motifs studied in ref. 28 (see Atoms 12-20 in Table 3). Interestingly, all atoms of the directed connectome contain at least one output node in the form of a node that has only incoming links.
E.coli Metabolic network
Next we consider the metabolic network of E.coli which is a part of a collection of 43 metabolic networks from29. For the E.coli metabolic network the MAP-configuration contains 11 non-trivial atoms (Table 4) which cover about 97% of the edges. The MAP configuration for the E.coli metabolic network has strong resemblance to a decomposition of the network into metabolic pathways. We find that the atoms in the MAP configuration of the metabolic network capture important features of metabolism and include motifs commonly used in biochemistry to describe metabolic pathways. For instance cycles (Atoms 3 and 9 in Table 4) and long bidirectional paths (Atom 10 in Table 4) are known to play an crucial role in regulating core metabolic pathways such as Gluconeogenesis30 and substrate cycles31. We also find atoms that can be interpreted as irreversible pathways (Atoms 2 and 6 in Table 4) where one metabolite is transformed into another in a controlled step-way fashion and atoms that can be interpreted as breaking down complex molecules into simpler ones which are then in turn combined to construct other larger molecules (Atoms 4 and 8 in Table 4). We also analysed the remaining 42 metabolic networks in the data set29 and found that the method finds very similar atoms across the networks in the data set (see Supplementary Note 5).
C.elegans neuronal network of synaptic connections
Finally, we consider the network of synaptic connections in the adult hermaphrodite worms C. elegans32. This network has 454 nodes and 4,841 directed edges. The network corresponding to male adults and the undirected networks representing gap junctions are discussed in Supplementary Note 5. The MAP configuration for the synaptic network 24 nontrivial atoms that cover approximately 70% of edges. We find that the MAP configuration contains a large number of bi-fan motifs (Atom 4 in Table 5) and multiple motifs that correspond to various combinations of feed–forward–loops (Atoms 6,7,8 and 10 in Table 5) indicating that feed–forward–loops usually associated to neuronal networks are organized into larger atoms in network. In addition to these the MAP configuration also contains multiple atoms resulting from symmetric combinations of the triangular connection pattern (A ↔ B, C → A, C → B) i.e. atoms 12, 14, 18 and 19 in Table 5. The combinations we find in general contain three or more smaller motifs in specific combinations suggesting the presence of motifs than go beyond pairwise combinations of lower order motifs as studied in28. Finally, we also observe atoms containing chains and cycles of bidirectional edges such as the atoms 1, 2, 15, 17, 20 and 21 in Table 5 and directed cliques. We also find the same general classes of atoms in the MAP configuration of the network of the male C.elegans for which the results can be found in Supplementary Note 5.
link

