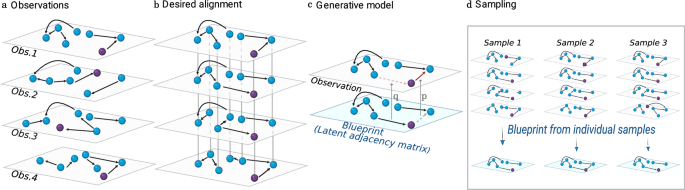
Probabilistic formulation of the problem of aligning multiple networks
Consider K network observations, with N nodes each and adjacency matrices {Ak; , k = 1, …, K }. We consider networks that are directed and with binary edges (that is, we just consider the presence or absence of connection between two nodes). Our hypothesis is that the observed networks are topologically similar, which allows us to map each node in a network to another node in each of the other networks. The goal is then to find the most plausible mapping of nodes across networks.
We follow a probabilistic approach and surmise that there is a latent underlying blueprint L, such that each network observation has been generated from that blueprint. Then, the network alignment problem becomes a problem of finding the blueprint L, and the permutations {π k, k = 1, …, K } that map nodes in each network to nodes in the blueprint. This formulation naturally allows the simultaneous alignment of several networks onto a single blueprint (Fig. 1).
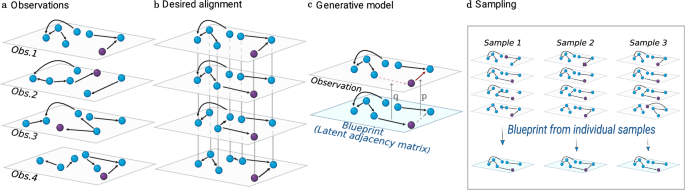
a Our data consists of observations of topologically similar networks with unknown node identities. b Our objective is to align the networks, that is, to find the permutation of node identities (illustrated as a rearrangement in the plane) in each network, such that each node is mapped to its counterpart in the other networks, and edges are as similar as possible across networks. To ease visual tracking, we color in purple a node that has the same hidden identity in all networks. c Generative model. We assume there exists an underlying blueprint from which observations are generated by copying edges and non-edges, with copying error probabilities q and p, respectively. d Our probabilistic approach allows us not only to formulate the network alignment problem in terms of finding the most plausible alignment, but also to sample over the space of possible alignments (that is, permutations of node identities) and corresponding blueprints. This allows us to assign a probability to each individual node mapping, and turns out to be critical to recover ground truth alignments in noisy networks (see text and Fig. 2).
Formally, we consider a blueprint L with binary edges (Lij ∈ {0, 1}) from which networks are copied with errors. Each entry Aij in an observed network is assumed to be independently generated from Lij with a copying error probability that depends on Lij. Edges in the blueprint (Lij = 1) are copied with error probability q, whereas non-edges (Lij = 0) are copied with error probability p (see Supplementary Material for the model with uniform copy error probability q = p). This approach shares some features with the approach in Ref. 22, where the goal is to align a noisy network copy with the original network (the blueprint in our case) using a similar probabilistic framework. The crucial difference lies in that, in our case, the blueprint is unknown, so that we need to infer it. This difference is precisely what makes our approach symmetric (the alignment of two networks does not require that we choose one as reference) and what allows us to align multiple networks at the same time without the need to choose an arbitrary blueprint from the set of observed networks.
For simplicity, we consider first the case in which we have a single observation. Given Lij, q, and p, the probability that we observe an edge (Aij = 1) or a non-edge (Aij = 0) is then:
$$p\, ({A}_{ij}=1| {L}_{ij},q,p)={p}^{(1-{L}_{ij})}{\left(1-q\right)}^{{L}_{ij}}$$
(1)
$$p\, ({A}_{ij}=0| {L}_{ij},q,p)={q}^{{L}_{ij}}{\left(1-p\right)}^{(1-{L}_{ij})}.$$
(2)
Since the probability of observing each edge is conditionally independent on the others, the total likelihood of observing an adjacency matrix A given a latent blueprint L, q, and p factorizes. Additionally, note that the likelihood depends on the unknown mapping of nodes in the observed network into nodes of the blueprint, which is given by the permutation π of the nodes in the observed network. With this, the likelihood is
$$p({{{\bf{A}}}}| {{{\bf{L}}}},q,p,\pi )={\prod}_{ij}p({A}_{ij}| {{{\bf{L}}}},q,p,\pi )={q}^{{o}_{10}}{p}^{{o}_{01}}{\left(1-q\right)}^{{o}_{11}}{\left(1-p\right)}^{{o}_{00}},$$
(3)
where the edge overlaps between the blueprint and the observations are \({o}_{XY}=\sum {\delta }_{{L}_{\pi (i)\pi (j)},X}{\delta }_{{A}_{ij},Y}\). For example, o01 is the number of entries that are 0 in L and 1 in A, for mapping π.
To obtain the posterior distribution over all model parameters, we use Bayes rule
$$p({{{\bf{L}}}},\pi,q,p| {{{\bf{A}}}})=\frac{p({{{\bf{A}}}}| {{{\bf{L}}}},q,p,\pi )\,p({{{\bf{L}}}},q,p,\pi )}{p({{{\bf{A}}}})},$$
(4)
where p(L, q, p, π) is the prior over model parameters. To obtain our desired posterior p(L, π∣A), we consider a beta prior over p and q, and integrate over these variables, which yields (“Methods”)
$$p({{{\bf{L}}}},\pi | {{{\bf{A}}}})\propto \frac{\Gamma ({o}_{11}+{\beta }_{q})\Gamma \left({o}_{10}+{\alpha }_{q}\right)}{\Gamma \left({n}_{1}+{\alpha }_{q}+{\beta }_{q}\right)}\frac{\Gamma ({o}_{00}+{\beta }_{p})\Gamma \left({o}_{01}+{\alpha }_{p}\right)}{\Gamma \left({n}_{0}+{\alpha }_{p}+{\beta }_{p}\right)},$$
(5)
where \({n}_{1}={\sum }_{i}{\delta }_{{L}_{ij},1}\), \({n}_{0}={\sum }_{i}{\delta }_{{L}_{ij},0}\) are the number of edges and non-edges in the latent blueprint L, respectively, and (αq, βq) and (αp, βp) are the hyper-parameters of the prior distributions for q and p, respectively (Methods). Note that one expects βq/p > αq/p ≥ 1 so that priors favor small copy error probabilities, although the effect of the prior becomes negligible for even modest edge overlaps.
In the case in which our observation comprises K networks, Eq. (5) generalizes to (“Methods”)
$$p({{{\bf{L}}}},\{{\pi }^{k}\}| \{{{{{\bf{A}}}}}^{k}\})\propto \frac{\Gamma ({O}_{11}+{\beta }_{q})\Gamma \left({O}_{10}+{\alpha }_{q}\right)}{\Gamma \left(K{n}_{1}+{\alpha }_{q}+{\beta }_{q}\right)}\frac{\Gamma ({O}_{00}+{\beta }_{p})\Gamma \left({O}_{01}+{\alpha }_{p}\right)}{\Gamma \left(K{n}_{0}+{\alpha }_{p}+{\beta }_{p}\right)},$$
(6)
where \({O}_{XY}={\sum }_{k}{o}_{XY}^{k}\), so that the posterior depends on the overall overlap of edges and non-edges of all the networks with the blueprint. Note that because β > α, the (L, {πk}) that maximize the posterior are the ones that maximize O11 and O00, and minimize O10 and O01. Also note that for a fixed permutation choice {πk}, the L⋆ that maximizes the posterior is such that \({L}_{ij}^{\star }\) is equal to the majority of all \({A}_{{\tilde{\pi }}^{k}(i),{\tilde{\pi }}^{k}(j)}^{k}\), where \({\tilde{\pi }}^{k}(i)\) is the inverse of πk(i), so that \({\pi }^{k}({\tilde{\pi }}^{k}(i))=i\) (“Methods”).
Note that, in the limit p → 0 or, equivalently, βp → ∞, our generative model is equivalent to generating networks from the blueprint by just sampling on the edges in the blueprint (that is Lij = 1). The limits of recovery of node identities and group labels of these nodes in terms of the sampling probability have been extensively studied for correlated pairs of random graphs31,32,33 and for correlated pairs of graphs with group structure34. In this probabilistic framework, the best alignment corresponds to the blueprint-permutation pair that maximizes the posterior probability p(L, {πk}∣{Ak}). In the limit p → 0, the posterior reduces to p(L, {πk}∣{Ak}) ∝ Γ(O11 + βq)Γ(O10 + αq), so that the most plausible alignment corresponds to the blueprint-permutation pair that maximizes O11, that is, the total number of edges in the graph that overlap with the blueprint. Formally, for the alignment of a pair of networks with binary adjacency matrices (A, B), if we assume that one of the two networks is equal to the blueprint as in Ref. 22, then maximizing O11 is equivalent to maximizing the product ∑ij Aij Bπ(i)π(j) as in the Koopmans-Beckmann QAP formulation of the graph matching problem11, and, in general, to minimizing the Frobenius norm between two adjacency matrices with binary entries. When p ≠ 0, our approach considers the contribution of both aligned edges and aligned non-edges. Therefore, the most plausible alignment in our case does not necessarily coincide with that in the Koopmans-Beckmann QAP formulation of the network alignment problem.
This observation also highlights the fact that current structural pairwise approaches cannot be easily modified to allow for the alignment of multiple networks, since, in order to do that, we would have to decide which of the networks is the generating blueprint against which all other networks need to be aligned. Unfortunately, if all copies are subject to the same copy error, there is no reason to expect that one of the observations is closer to the blueprint than another a priori. In fact, even if we had that prior information, pairwise approaches would only consider independent alignments of each network against the selected blueprint, so that the information coming from the overlap between all other pairs of networks other than the blueprint would be lost, and we would have to resort to adding a method to assess consistency of pair alignments29,30. Our probabilistic approach naturally circumvents these issues.
Sampling the space of plausible alignments
By analogy to statistical mechanics, we can associate an “energy” \({{{\mathcal{H}}}}=-\log \left(p({{{\bf{L}}}},\{{\pi }^{k}\},\{{{{{\bf{A}}}}}^{k}\})\right)\) to each blueprint-permutation pair (L, {πk}), so that their posterior is written
$$p({{{\bf{L}}}},\{{\pi }^{k}\}| \{{{{{\bf{A}}}}}^{k}\})=\frac{\exp \left(-{{{\mathcal{H}}}}\right)}{p(\{{{{{\bf{A}}}}}^{k}\})},$$
(7)
and the energy \({{{\mathcal{H}}}}\) can be obtained directly from Eq. (6). In this interpretation, the best alignment is the blueprint-permutation pair that minimizes \({{{\mathcal{H}}}}\). Importantly, this equivalence enables us to sample over the space of alignments (L, {πk}) using Markov chain Monte Carlo35 (“Methods”; see Supplementary Material and Fig. fig:time_size for the algorithmic complexity of our tool). The sampling of equilibrium alignments from this space allows us to approach the network alignment problem in ways other than just finding the single best alignment.
In particular, by sampling alignments from the posterior distribution, we can estimate the probability p(πk(ik) = iL∣{Ak}) that node ik in network Ak is mapped to node iL in the blueprint as the fraction of sampled permutations in which πk(ik) = iL. Then, we can estimate the most probable mapping for each node πk,⋆(ik) as the one that maximizes p(π(ik) = iL∣{Ak}) (Methods). As we show below, in noisy observations, the most plausible alignment does not necessarily recover the ground truth mapping of nodes. This is because for nodes with few connections, copy error can introduce a degree of ambiguity and degeneration in the mappings of node identities. However, we find that the most probable mapping of each individual node recovers the ground truth for their mapping more reliably (Fig. 2). This is, in fact, an expected result from using a probabilistic approach: averages over the ensemble of possible alignments are more accurate at predicting hidden information (in this case, node mappings to the blueprint) than single point estimates (that is, the best alignment) (see for instance Ref. 36 for a discussion in the context of link prediction).
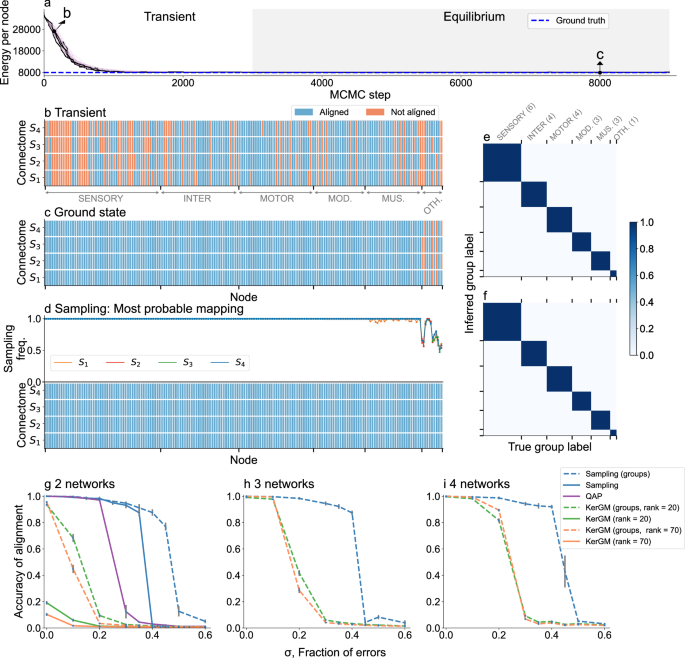
We consider C. elegans connectome A2 in Ref. 25 as a blueprint to generate noisy synthetic connectomes. a–d Alignment of 4 synthetic connectomes using group labels (neuron type) of each node. a Energy \({{{\mathcal{H}}}}\) as a function of time (Methods). Each line corresponds to a replica running at a different temperature. We sample alignments in the equilibrium zone (gray) at temperature T = 1 (black line). The blue dashed line shows the energy of the ground truth alignment. b–d. Alignment of individual nodes for each synthetic connectome S1, …, S4, ordered by neuron type. Blue nodes are correctly aligned, i.e., mapped to their ground truth identity (blue); red if misaligned (“Methods”). b Example of alignment in the transient regime—point b in (a). c. alignment with minimum energy (ground state)—point c in (a, d). Most probable mapping for each node sampled from the posterior distribution at equilibrium (text and Methods). The top panel shows the frequency with which each node (in each connectome) is assigned to its most probable mapping. e, f Node label inference. We assume that neuron types are unknown for some nodes in networks S2, S3, and S4 (total number for each type in parenthesis). We estimate the probability that a node has label X as the fraction of the sampled alignments in which that node is assigned that label (text and Methods). Each matrix element (X, Y) shows the probability that a node with unknown label Y is assigned label X. e Unlabeled nodes are the same across the synthetic connectomes S2−S4. f Unlabeled nodes are chosen at random in each of the connectome networks S2 − S4. See supplementary Figs. S5, S6 for their corresponding alignments. g–i Accuracy ar recovering the ground truth alignment for K noisy connectome copies with different fractions of errors, for three alignment methods: Fast QAP (QAP)38, multi-way (KerGM)23, and our sampling method (Sampling). g K = 2; h K = 3; i K = 4. Points show the mean accuracy for 10 sets of networks; error bars show the standard error of the mean.
Additionally, MCMC sampling allows us to leverage relevant information about the nodes. For instance, if we have access to node attributes such as group labels, we can constrain permutations to those in which only nodes with the same group labels can be mapped to the same node in the blueprint (Methods); or if we have information about the precise identity of some of the nodes (often called ‘anchors’ or ‘seeds’12; see Methods for details), our algorithm can be forced to only sample permutations that have a fixed mapping for these nodes.
Validation of the probabilistic approach on synthetic networks
We start by validating our approach on synthetic, but realistic, settings of increasing difficulty. Because network alignment is particularly relevant in the context of connectome analysis, we consider the connectome of the nematode C. elegans25,37 as our benchmark, and perform two initial sets of experiments: (i) the alignment of multiple identical networks in which node identities have been shuffled; (ii) the alignment of several noisy copies of the same network.
In the first experiment, we use multiple copies of the connectome network provided in ref. 11 (279 neurons), with node identities randomly shuffled in each copy. Because the networks are identical, the ground state of the energy in Eq. (7) coincides with the ground truth alignment, that is, with the desired mapping of node identities. However, the ground state is degenerate because any permutation of the nodes in the latent blueprint leaves the energy invariant. To break this symmetry, which greatly complicates the search in the space of possible alignments, we find that it is more efficient to first align two networks (chosen at random), and then add the remaining networks one by one to eventually obtain the desired global alignment of all the networks (Supplementary Fig. S2). We find that this approach enables us to perfectly align as many networks as desired, always converging to the ground state.
Except for the fact that we consider an arbitrary number of networks, as opposed to just two, the previous experiment is the standard validation test in the literature on network alignment and graph matching. In the second experiment, we move to a more realistic and challenging scenario. In particular, we use the adult A2 connectome of C. elega ns from ref. 25 as a blueprint (224 neurons and 2,186 connections), and generate four noisy synthetic connectomes by swapping 327 edges with non-edges from the original adjacency matrix.
Using a maximum likelihood estimate, this process would be equivalent to generating connectomes using our copying mechanism with q ≈ 0.15 and p ≈ 0.007.
Since the four observations are not identical, now the global ground state may not correspond to that of the ground truth. Additionally, if due to observational noise there are neurons with few connections, new degeneracies or quasi-degeneracies may appear. Therefore, the energy landscape is likely to become rougher in this realistic scenario. Indeed, we find that, whereas alignment is still possible and most nodes are correctly matched, the MCMC finds alignments with lower energy than the ground truth, indicating that the exact ground truth has become undiscoverable. (Since in this experiment the synthetic networks are generated by the generative model underlying our approach, our approach is Bayes optimal and no other method would, in general, be able to identify the ground truth alignment in this case.)
Given the added difficulty of this scenario, we take advantage of the fact that network datasets often contain useful information about nodes, such as group labels; in the specific case of connectomes, one could typically have information about neuron types (depending on their neuroblast lineage). Therefore, we can restrict our Markov chain to permutations where each neuron is mapped to a neuron of the same type in the blueprint. This drastically reduces the space of possible alignment permutations, and allows us to sample the alignment space more thoroughly and efficiently (Fig. 2). (Alternatively, we can apply the same successive alignment strategy as before, wherein we start by aligning two networks add layers successively; Supplementary Fig. S3.) Our results confirm that there exist multiple alignment permutations (and the corresponding blueprints) that have energies lower than that of the ground truth alignment (Fig. 2a, c).
This observation is key for real-world scenarios, and shows that the ground-state alignment should not be taken as the best estimate for the identity of individual nodes. Rather, as previously discussed, for individual nodes one should rather use the mapping πk,⋆(ik) that maximizes the probability p(πk(ik) = iL∣{Ak}) that node ik in network Ak is mapped to node iL in the blueprint (Methods). As we show in Fig. 2d, the most probable mapping for each node is more accurate at recovering the ground truth node identities. In fact, we find that some neurons are easy to align, so that the majority of sampled alignments have the same node mappings for those neurons; but there are other neurons, typically with few connections (see Supplementary Fig. S4), for which there are several possible alignments and only through averaging can we recover the underlying node identity.
To benchmark the performance of our approach, we compare it to other methods for network alignment: the Fast QAP11,38 and the multi-way Kernel Graph Matching (KerGM)23. Fast QAP is a method to align pairs of networks and offers the possibility to use anchors in the alignment but does not allow to use node attributes such as group labels; KerGM can align simultaneously multiple networks and also incorporate node attributes in the alignment process (see Methods for details). For a systematic comparison, we generate sets of K ∈ {2, 3, 4} noisy copies of the A2 connectome by swapping a fraction σ of the edges in the network with non-edges, with σ ∈ [0, 0.6]. For K = 2 we compare our approach without using group labels to the Fast QAP and KerGM, and using group labels against KerGM (Fig. 2g; Supplementary Table S1). For K = 3, 4, we compare our approach to KerGM, using group labels as node attributes in both cases (Fig. 2h, i; Supplementary Table S1; Methods). Our approach outperforms both QAP and KerGM, since it is able to recover accurate alignments for larger fractions of errors. Importantly, our approach has superior accuracy even when we reduce the number of sampled alignments with run-times comparable to those of KerGM (Supplementary Table S1).
Finally, to further showcase the potential uses of our approach, we note that, in some real-world scenarios, node group labels are only known for some of the nodes. For instance, in large neuronal connectomes27,28,39,40 neuron type is not always straightforward to establish, and some neurons are left without annotation. In such cases, our probabilistic approach can be used to infer the labels of unannotated nodes. Specifically, for each alignment sampled from the posterior, each node ik in network k is mapped to a labeled node in the blueprint iL = πk(ik); and a node ik in network k with unknown group label g(ik) is automatically assigned the label of the node in the blueprint to which it is mapped g(ik) = g(iL). Therefore, by sampling alignments from the posterior, we can estimate the probability p(g(ik) = g) that an unannotated node has label g as the fraction of sampled alignments in which g(ik) = g. To assess the performance of our approach in this task, we consider the same noisy connectome networks as in Fig. 2a–d, and we assume that one of the networks is fully annotated, but the others have a fraction of unlabeled nodes. In Fig. 2e, f, we show that our approach is able to recover ground truth annotation for all nodes with unknown labels, both when unlabeled nodes are randomly chosen in each observed connectome, and in the harder case in which unlabeled neurons are the same across connectomes (Supplementary Figs. S5, S6).
The probabilistic approach correctly aligns real networks
Having validated our method on synthetic networks for which the underlying assumptions are fulfilled by construction, we now explore its performance on sets of real networks, for which the assumptions may not be fulfilled. We consider three real network datasets (see Data for details): (i) four neural connectomes of the C. elegans nematode corresponding to four different developmental stages25; (ii) the left and right hemispheres of the brain of the larva of D. melanogaster26; (iii) the yearly e-mail communication networks within an academic institution for four consecutive years41. In each of these cases, we make use of whatever information is available from the nodes, and incorporate it to to the MCMC sampling to make it more efficient; we compare our results to those of Fast QAP and KerGM under the same conditions. In Table 1, we show a summary of the comparison, and in what follows we discuss each experiment in detail.
C. elegans connectome at different stages of development
We consider four connectomes of C. elegans at different stages of development25: two late larval samples (L2, L3) and two adult samples (A1, A2). In this dataset, each connectome comprises 224 neurons and each neuron is associated to one of six different neuron types, so we can use this information to constrain the search of alignment permutations. Yet, the alignment of these connectomes is challenging because the density of neuron-to-neuron connections increases throughout development, so that L2 and L3 connectomes are sparser than A1 and A2 connectomes (Supplementary Fig. S4). This implies that the network observations are not fully consistent with our assumption that they have been generated from the blueprint with the same copy error probability. Nonetheless, we find that the most probable mapping of each neuron (the mapping that maximizes the probability p(πk(ik) = iL∣{Ak}), obtained by averaging over alignments sampled in the equilibrium zone) recovers the ground truth identity for 94% of the neurons (Fig. 3a–b), compared to 56% identities for KerGM (Table 1). By contrast, as for the synthetic connectomes, the ground-state alignment is significantly further from the ground truth, as it misaligns 33.5% of the neurons due to the left-right symmetry in the data (an issue that is also apparent in the typical alignments obtained using KerGM; Supplementary Fig. S7). Therefore, in this case, it is critical to average over the ensemble of alignments in which different nodes in different connectomes are misaligned, to recover an alignment closer to the biological ground truth. Importantly, the advantage of sampling is already apparent even if we substantially reduce the number of samples and run times (Supplementary Fig. S7; Table 1).
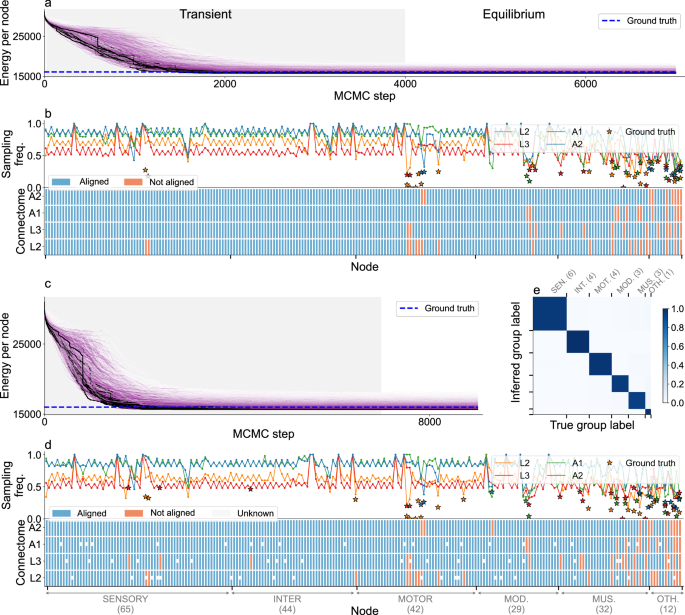
a, b Network alignment using group labels (neuron types). a Same as in Fig. 2a, b same as Fig. 2d.In this case the ground truth alignment (blue dashed line) is provided in ref. 25. Additionally, in (b) stars indicate the probability with which misaligned neurons are mapped to the ground truth identity.c–e Network alignment with some nodes with missing group labels. We assume that, for some nodes chosen at random in networks L2, L3 an A1, we do not have information about their group label (neuron type), while neurons in A2 are fully annotated. c, d same as a, b respectively.In the bottom panel of (d) nodes whose group label is assumed to be unknown are indicated by middle gray rectangles. e Probability matrix for the inferred group labels of the selected nodes, as in 2e, f. See Supplementary Figs. S9–S11 for results in slightly different experimental conditions.
To further benchmark the performance of our approach, we also looked at the alignment of two adult connectomes (A1, A2), in which case we can also compare to Fast QAP. In general, aligning pairs of networks happens to be a harder problem (because less information is available about nodes and their connections) but, again, the alignment obtained using our approach is better than those obtained using any of the other methods (Table 1; Supplementary Fig. S8). However, in contrast to the alignment of four connectomes, the most probable mapping does not improve over the ground state (Supplementary Fig. S8). This is because for only a pair of networks the alignment landscape is dominated by one deep minimum.
Finally, we investigate the ability of our approach to infer unknown group labels (that is, neuron types), assuming that A2 is fully annotated and that L2, L3, and A1 have 15% of the nodes without annotation (Fig. 3c–e). This is a harder problem, since unannotated nodes are unconstrained by group label and the number of possible permutations is, thus, larger. Nonetheless, we find that, again by sampling from the posterior distribution of alignments, the most probable mapping of each neuron recovers the ground truth identity in 92.8% of cases, and that our approach is able to fully recover unknown labels. Qualitatively, these results are robust to changes in the selection of the fully annotated connectome and the unannotated nodes, and to other variations in the experiment (Supplementary Figs. S9–S11).
Left and right brain hemispheres for the Drosophila melanogaster larva
Next, we consider the connectome for the full brain of the larva of the Drosophila melanogaster fly26. Neurons in the fly brain can be classified into hemispheres that are assumed to be mirror copies of one another, which is consistent with our hypothesis of an underlying blueprint. Our goal is thus to use our probabilistic approach to align the two hemispheres and show that our method is valid to align connectomes comprising over a thousand neurons. For this task, we constrain the search space of permutations by using two pieces of information (Data): (i) the neuroblast lineage of each neuron; and (ii) a set of 292 anchors, that is, pairs of neurons for which the precise mapping is well established (also called “seeds“ in the literature12). In Fig. 4, we show that the most probable mapping of each neuron recovers the ground truth mapping in 79.2% of the cases (or 78.6% if we consider a smaller set of alignments). This is higher than just looking at the ground state (77.1%), the best alignment we obtained using a seeded QAP approach12 (76.9%), and also the mean accuracy obtained using KerGM (69.5 %) (Table 1 and Supplementary Fig. S12).
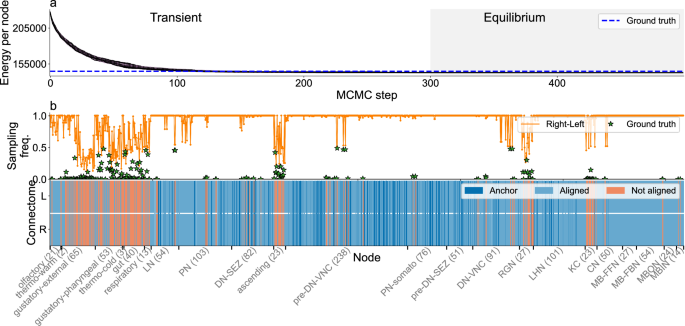
a Same as in Fig. 2a, b same as Fig. 2d. In this case the ground truth alignment (blue dashed line) is provided in ref. 26. In the top panel of (b) stars indicate the probability with which misaligned neurons are mapped to the ground truth identity. In the bottom panel apart from the aligned and misaligned nodes, there are nodes set as anchors (dark blue) as indicated in the dataset (292 out of 1235 in each hemisphere).
Note also that neither anchors nor mismatched neurons are homogeneously distributed. For example, some groups are hard to align because the topological overlap between connections of ground truth pairs and those with other neurons of the same type is often large, such as for Kenyon cells (KC; neurons in the mushroom body of the brain) or gustatory external neurons (Supplementary Figs. S13, S14). For KC neurons misalignment between neurons is to be expected, since these neurons establish many connections following a random pattern with well defined pre-synaptic and post-synaptic partners, and therefore they are topologically hard to distinguish42. Nonetheless, this is not necessarily the case for other neuron types such as gustatory neurons, for which there are clear topological partners. For some of these misaligned neuron pairs, we have checked that our tool finds the topologically optimal solution that does not correspond to the biological ground truth, suggesting that either the annotation is incorrect or that there is further biological information the we would need to incorporate into the tool to be able to recover the biological ground truth mapping.
E-mail communication networks
As a final case study, we consider a completely different network representing the emails exchanged between individuals within an academic institution during four consecutive years (2007-2010)41. For each year, we define a directed network where the nodes are individuals and links represent the existence of stable e-mail communication between them during the years we consider (Data). In particular, we consider two cases of stable yearly communication between individuals that result in two networks of 170 nodes (Aij = 1 if i sends at least 25 emails to j in a year) and 356 nodes (Aij = 1 if i sends at least 12 emails to j in a year), respectively. E-mail communication between two individuals can vary significantly from year to year41, which makes this a harder problem and serves as a benchmark for networks with large copy error.
The data includes the organizational unit to which each individual is affiliated, so we use this information to constrain the possible alignment permutations. In Fig. 5, we show that for both networks most of the alignment permutations we sample have an energy which is lower than that of the ground truth alignment. As in the case of C. elegans, we find that sampling over alignments increases significantly the ability to recover ground truth alignment. Using sampling we recover the ground truth identity for 95.9% and 96.1% of the nodes for the 170 and 356-node network, respectively (95.1% and 94% if we use a reduced sampling strategy), which is substantially higher than looking at the ground state alignment (92.8% and 89.3%) or using KerGM (average 80% and 59%) (Table 1 and Supplementary Fig. S15).
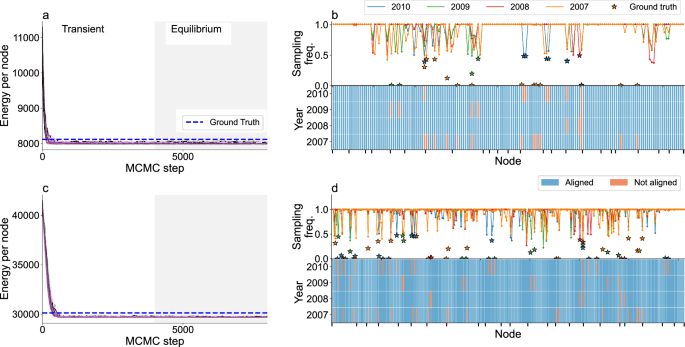
Multiple network alignment using the information about the group label (organizational unit) of each node, for different constrained set of users, (a, b) for 170 users and (c, d) for 356 users in 23 organizational units (see Data). a, c Same as in Fig. 2a, b, d same as Fig. 2d. In the top panels of (b, d) stars indicate the probability with which misaligned neurons are mapped to the ground truth identity.
link